Cell-state transition can reveal additional information from single-cell ribonucleic acid (RNA)-sequencing data in time-resolved biological phenomena. However, most of the current methods are based on the time derivative of the gene expression state, which restricts them to the short-term evolution of cell states. Researchers at Fudan University have developed single-cell State Transition Across-samples of RNA-seq data (scSTAR), which overcomes this limitation by constructing a paired-cell projection between biological conditions with an arbitrary time span by maximizing the covariance between two feature spaces using partial least square and minimum squared error methods. In mouse ageing data, the response to stress in CD4+ memory T cell subtypes was found to be associated with ageing. A novel Treg subtype characterized by mTORC activation was identified to be associated with antitumour immune suppression, which was confirmed by immunofluorescence microscopy and survival analysis in 11 cancers from The Cancer Genome Atlas Program. On melanoma data, scSTAR improved immunotherapy-response prediction accuracy from 0.8 to 0.96.
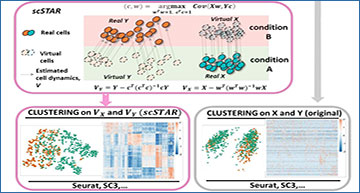
scSTAR schematic diagram and preliminary evaluation on scRNA-seq data
(a) The flow diagram of estimating single cell state transfer information, results of the clustering step are shown for the immunosenescence data4 5 . (b) scSTAR improves the immunotherapy outcome prediction accuracy. (c, d) scSTAR more clearly reveals the Peripheral Blood Mononuclear Cell (PBMC) discrimination between early and late stage lung cancer patients. (e, f) scSTAR better reconstructs dendritic cell transition trajectories from normal to tumor tissue.
Hao J, Zou J, Zhang J, Chen K, Wu D, Cao W, Shang G, Yang JYH, Wong-Lin K, Sun H, Zhang Z, Wang X, Chen W, Zou X. (2023) scSTAR reveals hidden heterogeneity with a real-virtual cell pair structure across conditions in single-cell RNA sequencing data. Brief Bioinform [Epub ahead of print]. [abstract]