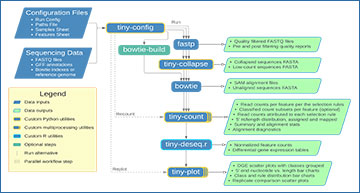
Researchers at Colorado State University have developed tiny-count, a highly flexible counting tool that allows for hierarchical classification and quantification of small RNA reads from high-throughput sequencing data. Selection rules can be used to filter reads by 5’ nucleotide, length, position of alignments in relation to reference features, and by the number of mismatches to reference sequences. tiny-count can quantify reads aligned to a genome or directly to small RNA or transcript sequences. With tiny-count, users can quantify a single class of small RNAs or multiple classes in parallel. tiny-count can resolve distinct classes of small RNAs, for example piRNAs and siRNAs, produced from the same locus. It can distinguish small RNAs variants, such as miRNAs and isomiRs, with single-nucleotide precision. tRNA, rRNA, and other RNA fragments can also be quantified. tiny-count can be run alone or as part of tinyRNA, a workflow that provides a basic all-in-one command line-based solution for small RNA-seq data analysis, with documentation and statistics generated at each step for accurate and reproducible results.
Availability – tiny-count and other tinyRNA tools are implemented in Python, C ++, Cython, and R, and the workflow is coordinated with CWL. tiny-count and tinyRNA are free and open-source software distributed under the GPLv3 license. tiny-count can be installed via Bioconda (https://anaconda.org/bioconda/tiny-count) and both tiny-count and tinyRNA documentation and software downloads are available at https://github.com/MontgomeryLab/tinyRNA.
Tate AJ, Brown KC, Montgomery TA. (2023) tiny-count: a counting tool for hierarchical classification and quantification of small RNA-seq reads with single-nucleotide precision. Bioinform Adv [Epub ahead of print]. [abstract]