In the world of modern biology, understanding the complexities of individual cells is essential. One powerful tool researchers use to study these intricacies is Single-Cell Ribonucleic Acid Sequencing (scRNA-seq). This high-throughput technique allows scientists to investigate the transcriptomes—the complete set of RNA transcripts—of single cells. However, analyzing scRNA-seq data is not without its challenges. The data is often high-dimensional, noisy, and sparse, making it difficult to accurately cluster and analyze.
The Need for Effective Clustering in scRNA-seq Data
Clustering is a method used to group similar cells based on their gene expression profiles. Effective clustering can reveal the diversity and heterogeneity of cells within a sample, which is crucial for understanding complex biological processes and diseases. However, traditional clustering algorithms struggle with the unique challenges presented by scRNA-seq data. This is where the Dual Correlation Reduction network-based Extreme Learning Machine (DCRELM) comes into play.
What is DCRELM?
DCRELM is a novel clustering algorithm specifically designed to handle the complexities of scRNA-seq data. Here’s how it works:
- Low-Dimensional Feature Extraction:
- DCRELM starts by transforming the high-dimensional scRNA-seq data into a low-dimensional space using an Extreme Learning Machine (ELM). This step reduces the complexity and makes the data more manageable.
- Dual View Enhancement:
- The ELM graph distortion module then generates a dual view of the features. This process enhances the robustness of the data by providing a more comprehensive representation of the underlying structures.
- Autoencoder Fusion:
- Next, the autoencoder fusion module learns both the attributes and structural information of the features. By merging these two types of information, it creates consistent latent representations that better capture the essence of the data.
- Noise and Redundancy Reduction:
- The dual information reduction network filters out redundant information and noise from the latent representations. This step ensures that the final data used for clustering is clean and precise.
- Triplet Self-Supervised Learning:
- Finally, a triplet self-supervised learning mechanism is employed to further refine the clustering performance. This mechanism improves the accuracy of the clusters by learning from the data itself without the need for additional labeled information.
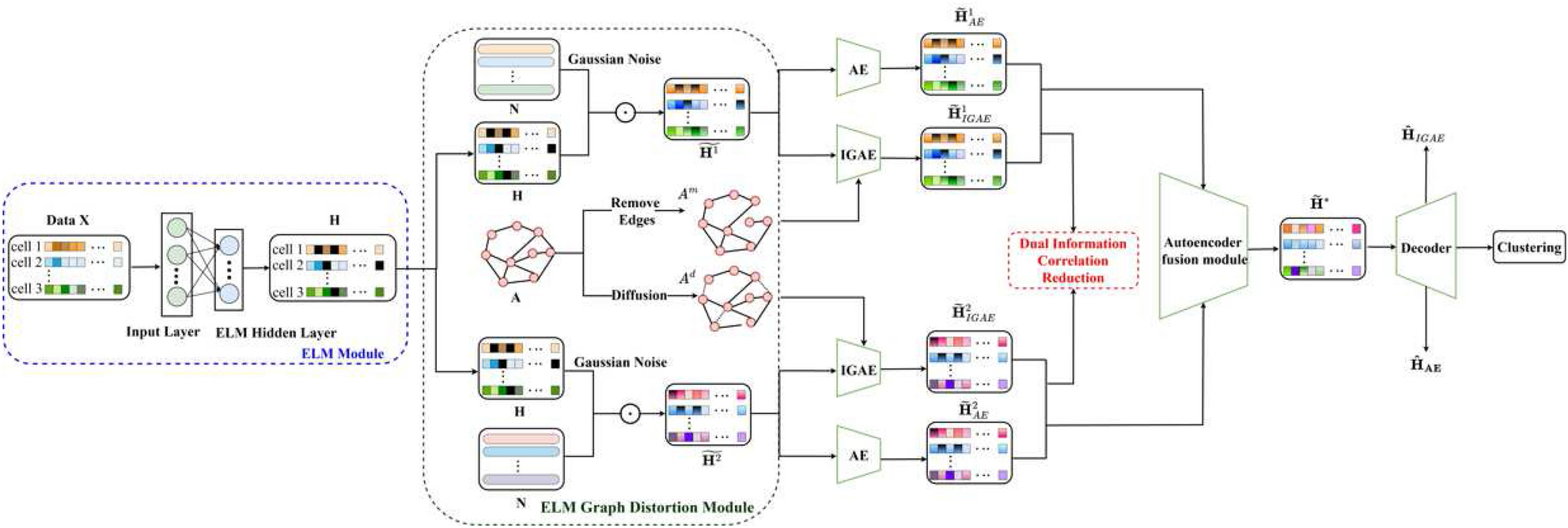
Framework diagram of the DCRELM
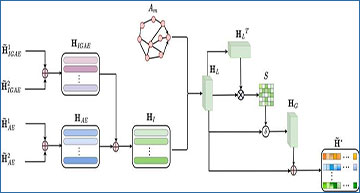
The cell-gene expression matrix X from the scRNA-seq data was selected as the input matrix. The ELM maps the original high-dimensional sparse X into a random mapping space to obtain a low-dimensional dense cell output matrix H. Using a siamese network framework, the attribute information of the cell output matrix H is enhanced to obtain ?~1 and ?~2, while the graph structure information of the cell adjacency matrix A is enhanced to obtain ?? and ??. Then, the fusion encoder in the autoencoder fusion module is employed to extract latent features ??1 and ??2 from ?~1 and ?~2, and the dual correlation information reduction network is utilized to remove noise and redundant feature information. High-quality cell-gene expression features are obtained by decoding the fusion module. The KL loss function of the triplet self-supervised strategy is minimized to improve clustering performance and effectively identify cell types.
Why is DCRELM Important?
DCRELM addresses several key challenges in scRNA-seq data analysis:
- Dimensionality Reduction: By transforming high-dimensional data into a low-dimensional space, DCRELM makes it easier to handle and analyze.
- Robustness: The dual view enhancement and autoencoder fusion steps create a more robust representation of the data, which is less sensitive to noise and variability.
- Precision: Noise and redundancy reduction ensure that the clustering results are precise and reliable.
- Self-Supervised Learning: The triplet self-supervised learning mechanism allows DCRELM to improve its performance by learning from the data itself.
Proven Performance
Extensive experiments have demonstrated that DCRELM performs exceptionally well in terms of clustering performance and robustness. It effectively reveals the heterogeneity and diversity of cells in scRNA-seq data, making it a valuable tool for researchers.
Conclusion
The DCRELM algorithm represents a significant advancement in the analysis of single-cell transcriptomes. By addressing the unique challenges of scRNA-seq data, DCRELM enables more accurate and robust clustering, providing deeper insights into the complexity of individual cells. This breakthrough has the potential to enhance our understanding of various biological processes and diseases, paving the way for new discoveries in the field of genomics.
Availability – The code is available at https://github.com/gaoqingyun-lucky/awesome-DCRELM.
Gao Q, Ai Q. (2024) DCRELM: dual correlation reduction network-based extreme learning machine for single-cell RNA-seq data clustering. Sci Rep 14(1):13541. [article]