RNA-sequencing technology provides an effective tool for understanding miRNA regulation in complex human diseases, including cancers. A large number of computational methods have been developed to make use of bulk and single-cell RNA-sequencing data to identify miRNA regulations at the resolution of multiple samples (i.e. group of cells or tissues). However, due to the heterogeneity of individual samples, there is a strong need to infer miRNA regulation specific to individual samples to uncover miRNA regulation at single-sample resolution level.
Researchers at Dali University have developed a framework, Scan, for scanning sample-specific miRNA regulation. Since a single network inference method or strategy cannot perform well for all types of new data, Scan incorporates 27 network inference methods and two strategies to infer tissue-specific or cell-specific miRNA regulation from bulk or single-cell RNA-sequencing data. Results on bulk and single-cell RNA-sequencing data demonstrate the effectiveness of Scan in inferring sample-specific miRNA regulation. Moreover, the researchers have found that incorporating priori information of miRNA targets can improve the accuracy of miRNA target prediction. In addition, Scan can contribute to the clustering cells/tissues and construction of cell/tissue correlation networks. Finally, the comparison results have shown that the performance of network inference methods is likely to be data-specific, and selecting optimal network inference methods is required for more accurate prediction of miRNA targets. The researchers have made Scan freely available to the public to help infer sample-specific miRNA regulation for new data, benchmark new network inference methods and deepen the understanding of miRNA regulation at the resolution of individual samples.
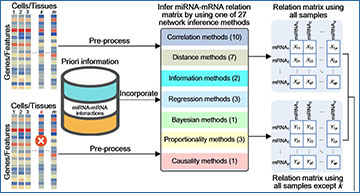
Schematic illustration of Scan
Given bulk or single-cell RNA-sequencing data with or without priori information of miRNA-mRNA interactions, Scan can apply 27 computational methods covering 7 types (correlation, distance, information, regression, bayesian, proportionality and causality) to construct miRNA-mRNA relation matrix. By using one of 27 computational methods, Scan constructs two miRNA-mRNA relation matrices (one for all samples and the other for all samples except k). For each sample (cell or tissue) k, Scan conducts sample-specific network inference from the two constructed miRNA-mRNA relation matrices to infer a miRNA regulatory network specific to the sample k. In total, Scan can identify m sample-specific miRNA regulatory networks across m samples (one network for one sample). In terms of accuracy and scalability, Scan further evaluates the constructed sample-specific miRNA regulatory networks.
Availability – Scan is released under the GPL-3.0 License, and is available at https://github.com/zhangjunpeng411/Scan/.
Zhang J, Liu L, Wei X, Zhao C, Luo Y, Li J, Le TD. (2023) Scanning sample-specific miRNA regulation from bulk and single-cell RNA-sequencing data. bioRXiv [online preprint]. [article]