Using single-cell RNA sequencing (scRNA-seq) data to diagnose disease is an effective technique in medical research. Several statistical methods have been developed for the classification of RNA sequencing (RNA-seq) data, including, for example, Poisson linear discriminant analysis (PLDA), negative binomial linear discriminant analysis (NBLDA), and zero-inflated Poisson logistic discriminant analysis (ZIPLDA). Nevertheless, few existing methods perform well for large sample scRNA-seq data, in particular when the distribution assumption is also violated.
Shenzhen University researchers have developed a deep learning classifier (scDLC) for large sample scRNA-seq data, based on the long short-term memory recurrent neural networks (LSTMs). Their new scDLC does not require a prior knowledge on the data distribution, but instead, it takes into account the dependency of the most outstanding feature genes in the LSTMs model. LSTMs is a special recurrent neural network, which can learn long-term dependencies of a sequence.
Simulation studies show that this new scDLC performs consistently better than the existing methods in a wide range of settings with large sample sizes. Four real scRNA-seq datasets are also analyzed, and they coincide with the simulation results that our new scDLC always performs the best.
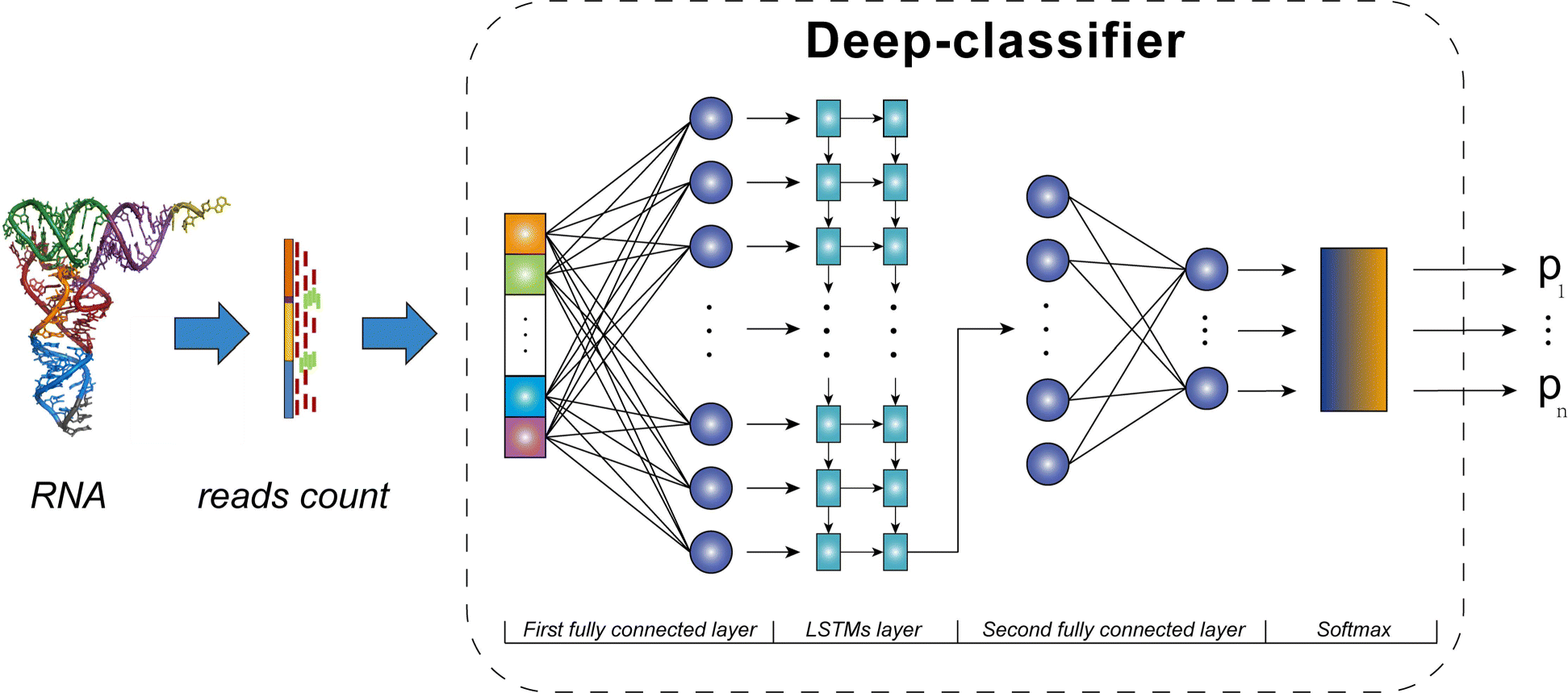
The framework of scDLC with three layers
The first layer and the third layer are two fully connected layers, and the middle layer is an LSTMs layer that consists of two long short-term memory network sub layers. A softmax layer is connected at the end to map the output of the classifier to a probability distribution
Availability – The code named “scDLC” is publicly available at https://github.com/scDLC-code/code.
Zhou Y, Peng M, Yang B, Tong T, Zhang B, Tang N. (2022) scDLC: a deep learning framework to classify large sample single-cell RNA-seq data. BMC Genomics 23(1):504. [article]